We utilize a combination of fieldwork, ecological data, genomics, and bioinformatics to study how organisms evolve across both geographic and genomic landscapes.
Avian Population Genomics and Genome Evolution
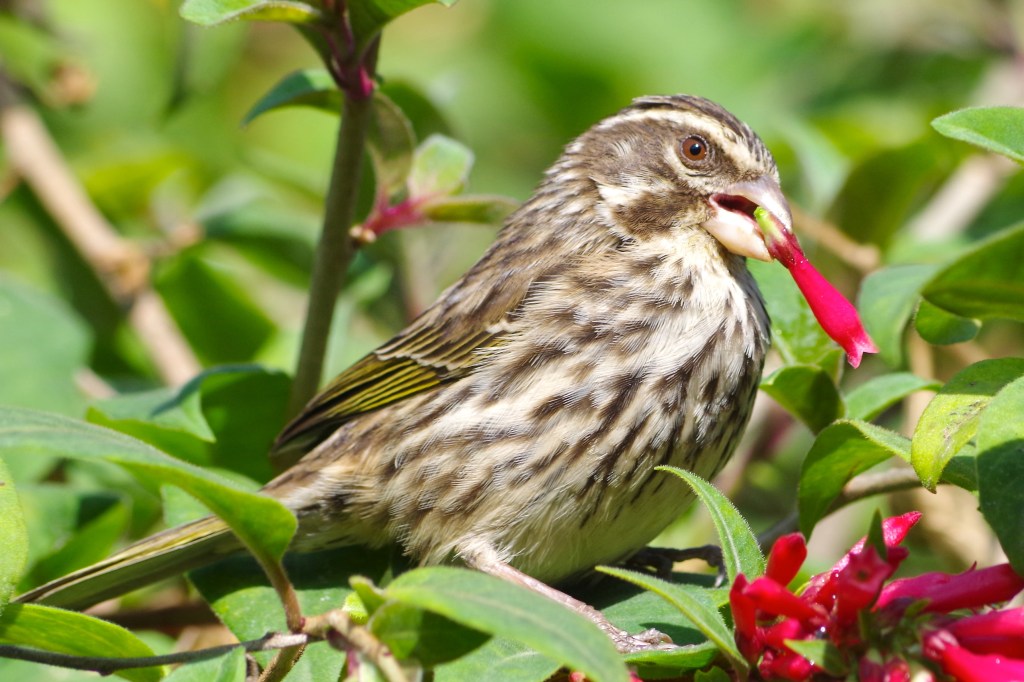
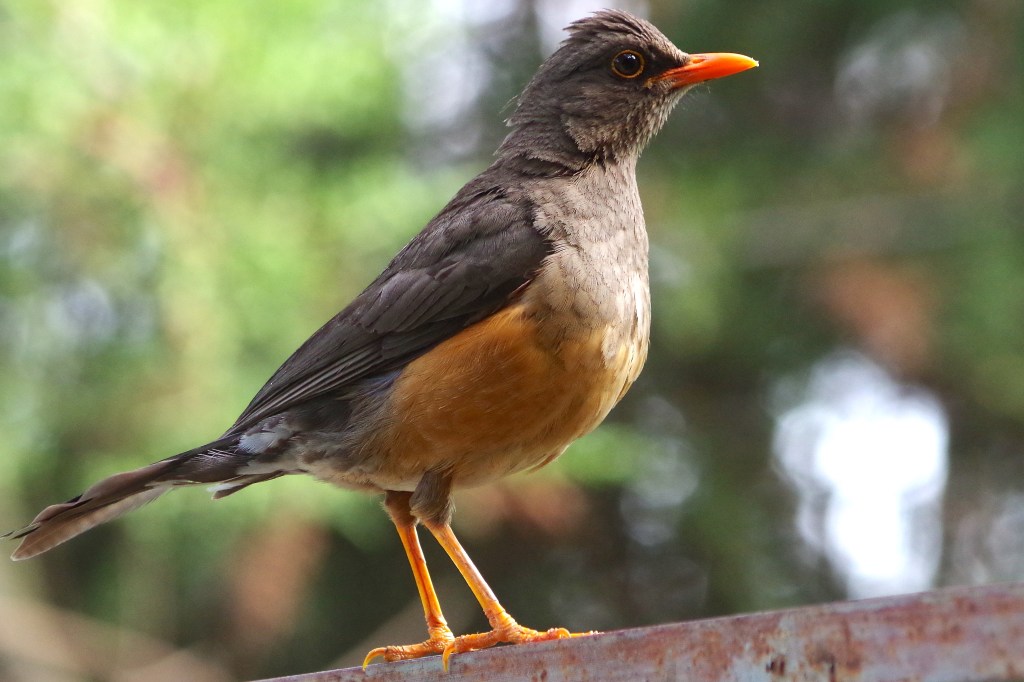
We are studying the population genomics and genome evolution of birds at multiple taxonomic scales (within populations, between populations, between closely related species) using multiple sequencing types, including long-read data, whole-genome resequencing Illumina data, and transcriptome data. Our goals are to better understand the evolutionary dynamics of diversification and to understand how structural variants (including indels, inversions, transposable elements, etc.) are evolving in genomes. Lastly, we want to better understand how structural variants impact genome evolution, molecular evolution, population genomics processes, and diversification.
Current Projects:
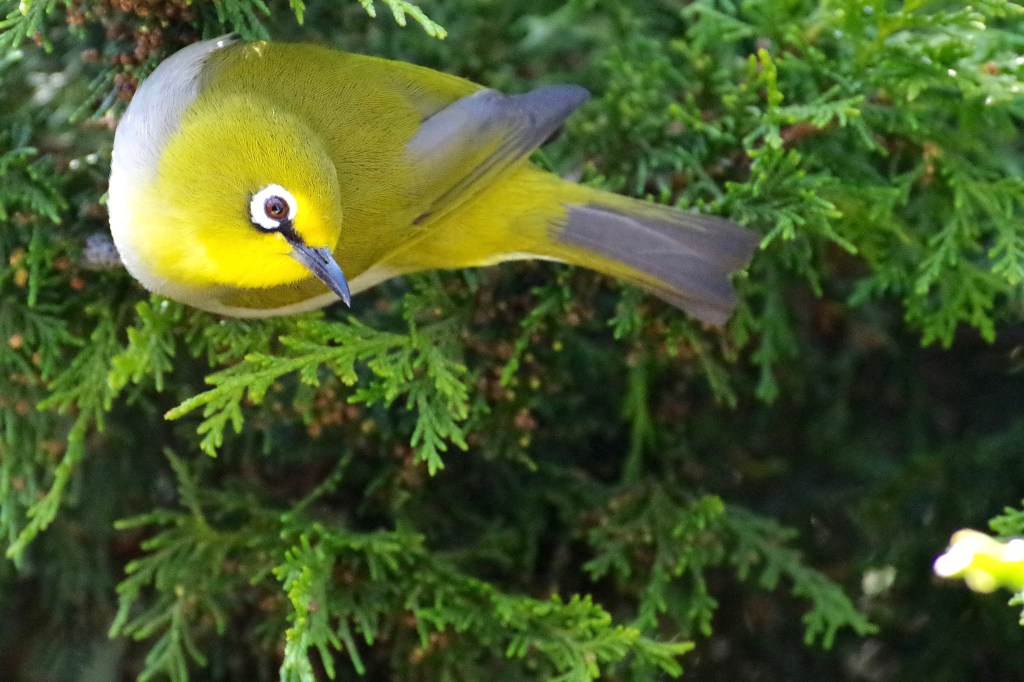
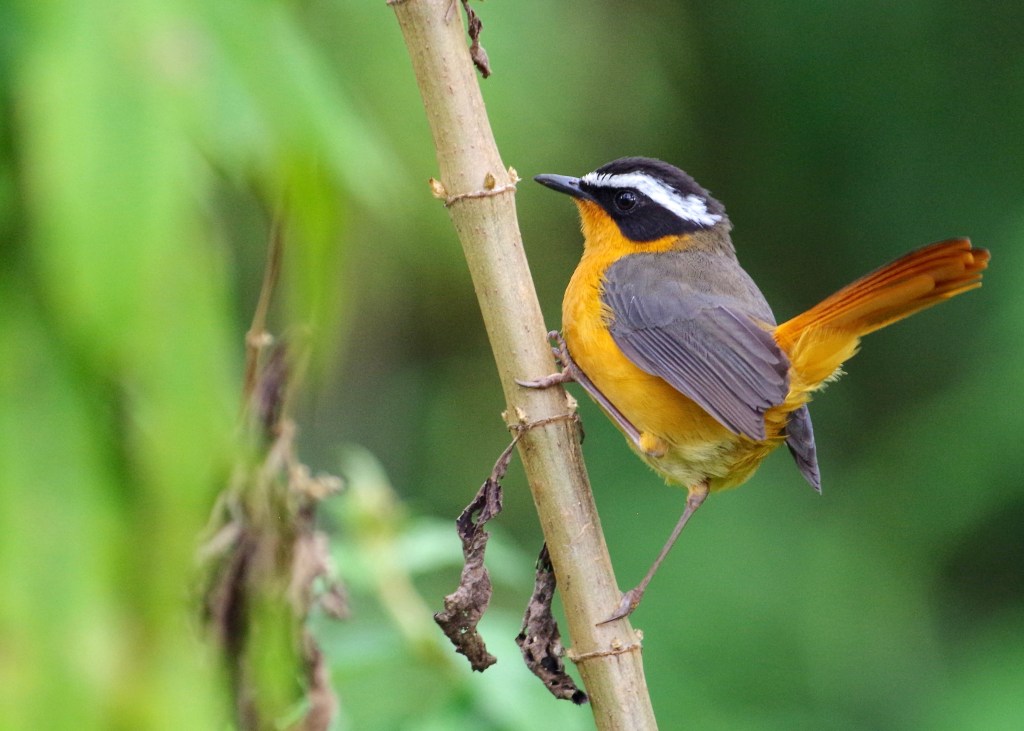
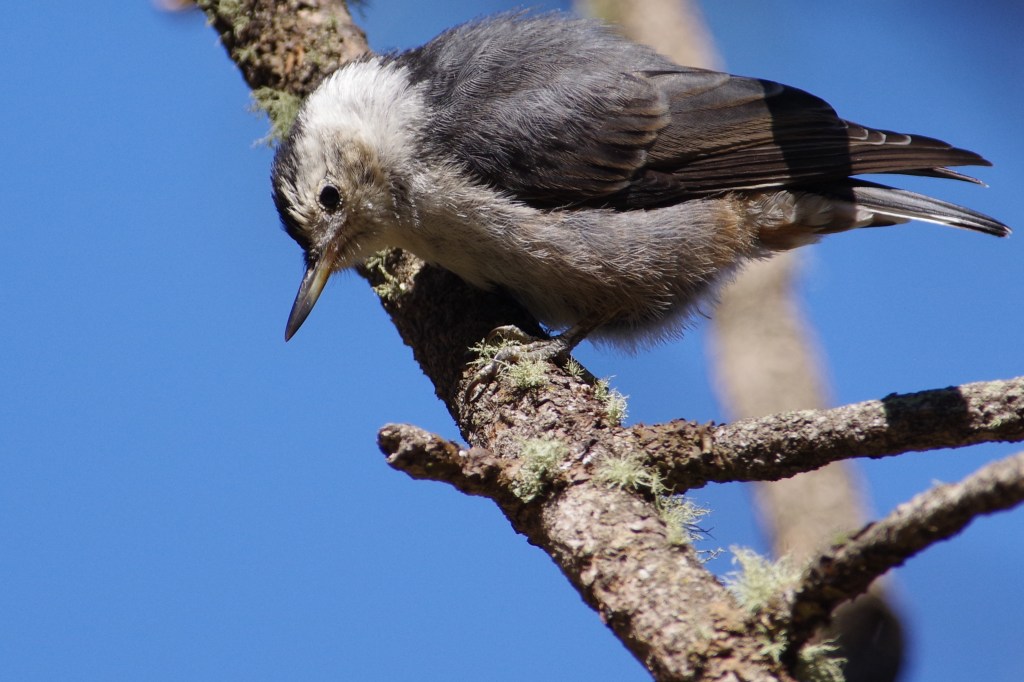
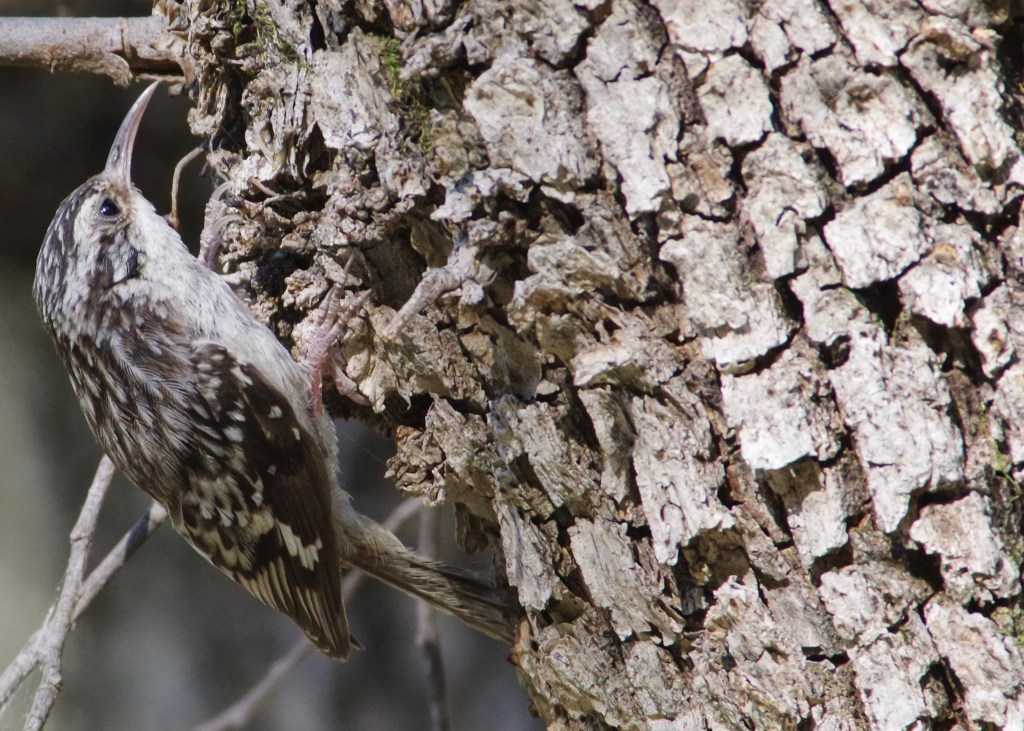
- Population genomics, landscape genomics, hybrid zone genomics, and phylogeography in several avian taxa (including species in the genera: Certhia, Contopus, Leiothlypis, Pachycephala, Setophaga, Sitta, Zosterops).
- Biodiversity genomics through time of Eastern African montane birds (animated video: https://www.youtube.com/watch?v=MfUO7oQc3zI).
- Population genomics of structural variants in multiple avian groups.
Recent Relevant Products:
- Manthey JD, Settlecowski AE, Meheretu Y, Behrends GJ, Bourgeois Y, Campillo LC, Boissinot S & Marks BD (2025) Temporal genomics reveals a century of genomic diversity shifts in a biodiversity hotspot avian assemblage. Genome Biology and Evolution evaf163. https://doi.org/10.1093/gbe/evaf163
- Gyllenhaal EF, Andersen MJ, Moyle RG & Manthey JD (2025) Island size shapes genomic diversity in a great speciator (Aves: Zosterops). Biology Letters. https://doi.org/10.1098/rsbl.2024.0692
- Behrends GJ, Meheretu Y & Manthey JD (2024) The Great Rift Valley is a greater biogeographic barrier than the Blue Nile Valley for six Ethiopian Highland passerines in the eastern Afromontane biodiversity hotspot. Ornithology. https://doi.org/10.1093/ornithology/ukae030
- Manthey JD & Spellman GM (2024) Recombination rate variation shapes genomic variability of phylogeographic structure in a widespread North American songbird (Aves: Certhia americana). Molecular Phylogenetics and Evolution. https://doi.org/10.1016/j.ympev.2024.108088
- Settlecowski AE, Marks BD & Manthey JD (2023) Library preparation method and DNA source influence endogenous DNA recovery from 100-year-old avian museum specimens. Ecology and Evolution. DOI: 10.1002/ece3.10407






Ant-Microbe Co-Evolution and Genome Evolution (currently NSF funded)
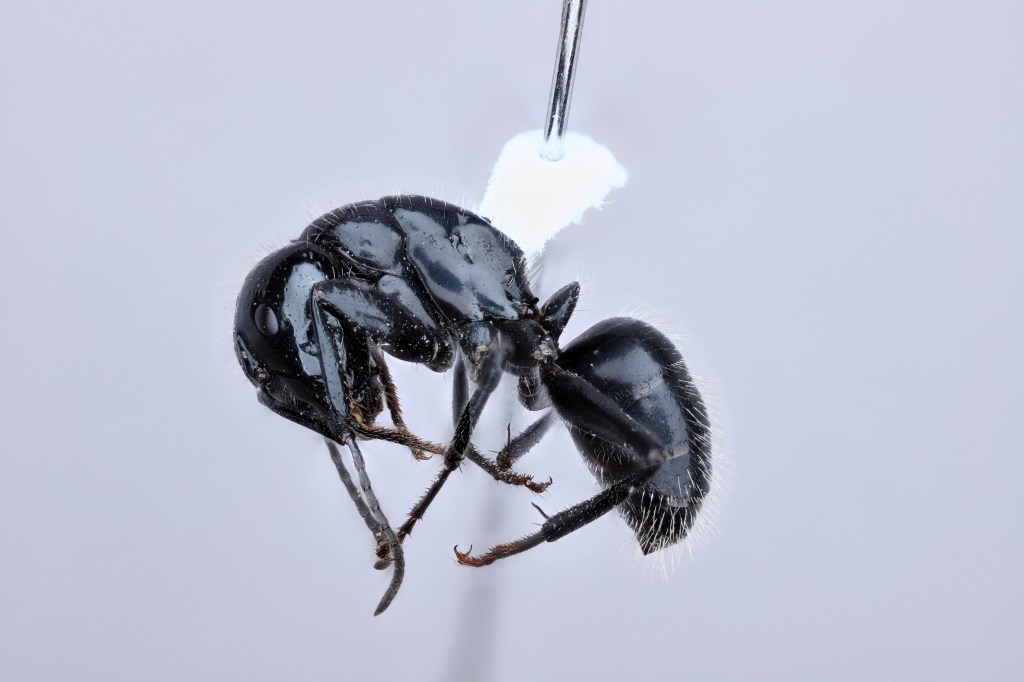
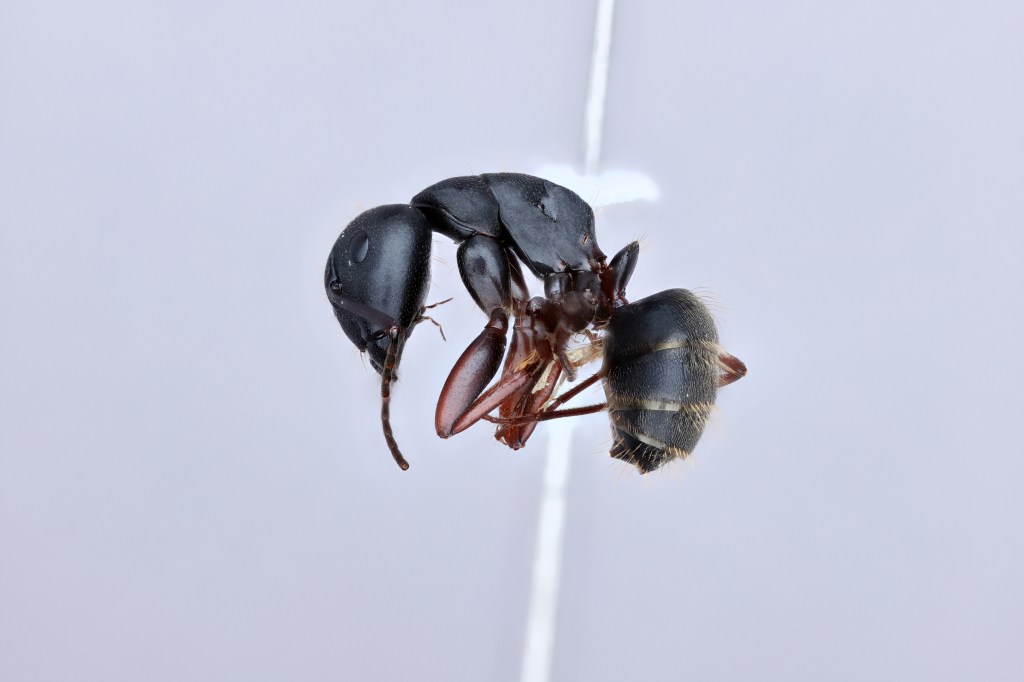
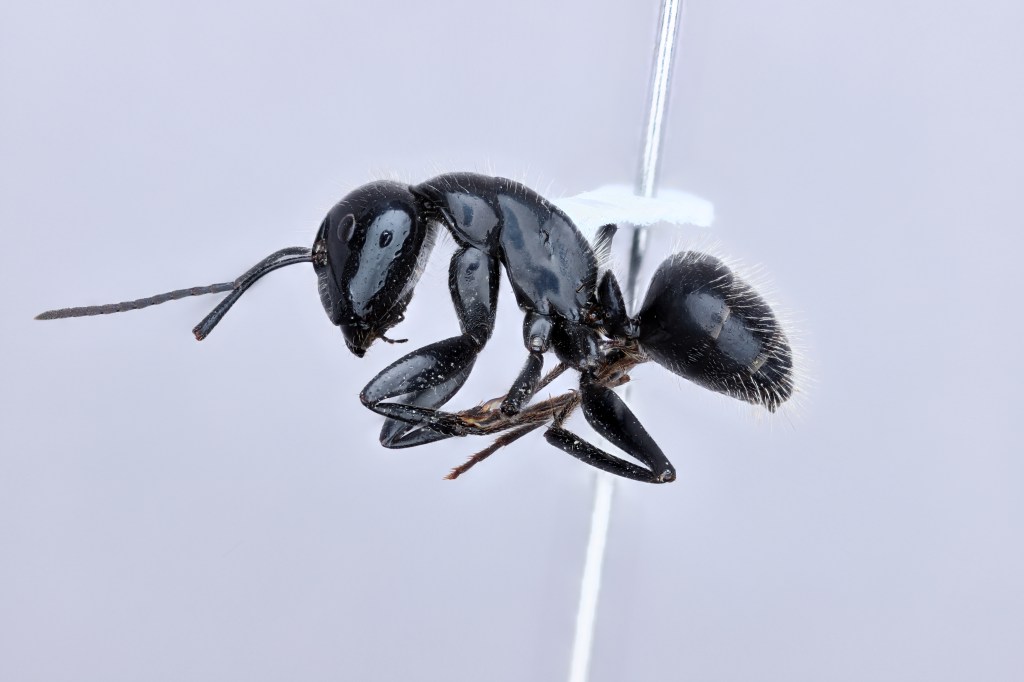
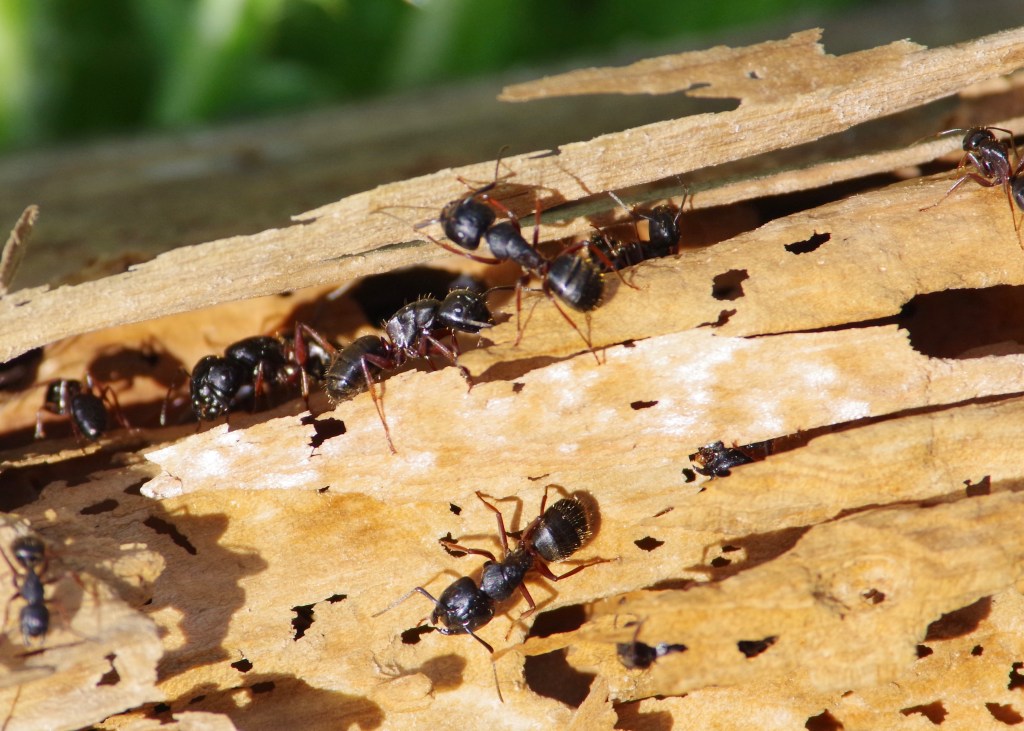
We are studying the codiversification and genome evolution of Camponotus carpenter ants and their Blochmannia endosymbionts using whole genome sequencing methods and ecological data. We are currently focusing on North American taxa.
Current Projects:
- Effects of host demography on endosymbiont genome evolution
- Landscape genomics of several North American Camponotus species
- Hybridization between Camponotus species
- Microbial community ecology in Camponotus hosts
- Population genomics of structural variants in carpenter ants (and some other invertebrates).
- Genome evolution in Blochmannia endosymbionts of carpenter ants
Relevant Products:
- Meadows BA, Emad M, Hruska JP, Silva J. Behrends GJ, Girón JC, & Manthey JD. 2023. Relatedness within colonies of three North American species of carpenter ants (Subgenus: Camponotus) and a comparison with relatedness estimates across Formicinae. Insectes Sociaux 70, 191-202.
- Manthey JD, Girón JC & Hruska JP. 2022. Impact of host demography and evolutionary history on endosymbiont molecular evolution: A test in carpenter ants (genus Camponotus) and their Blochmannia endosymbionts. Ecology and Evolution doi: 10.1002/ece3.9026





Conservation Genomics
We are always pursuing funding for conservation genomics projects relevant to local and regional stakeholders. Currently, we are working on conservation genomics of a couple species in Texas, including the Palo Duro Mouse (Peromyscus truei comanche) and American Black Bear (Ursus americanus). In addition to our research group, these projects are in collaboration with Caleb Phillips and Robert Bradley at TTU.
These projects generally consist of using whole-genome sequencing to address population and conservation genomics questions of interest, including assessment of regional genomic distinctiveness, connectivity among populations, and regional levels of genetic diversity.

