(Presented in order of joining the group)
Joe Manthey
- 2024-Present – Assoc. Professor Texas Tech
- 2018-2024 – Asst. Professor Texas Tech
- 2016-2017 – Postdoc NYU Abu Dhabi
- 2011-2015 – Ph.D. University of Kansas
- 2009-2011 – M.S. Black Hills State University
- 2004-2008 – B.S. Univ. of Wisconsin – Madison
Contact : joseph.manthey (at) ttu.edu
Google Scholar – GitHub

Ari Rice
- 2021-Present – Ph.D. Student Texas Tech
- 2020 – M.S. Villanova University
- 2016 – B.A. Lawrence University
Contact : arrice (at) ttu.edu
Google Scholar
I am interested in using genomic tools to study past and present gene flow between populations of birds living in sky island ecosystems. I am also generally interested in avian hybrid zones, investigating the ecological and landscape-related factors that make hybridization possible between certain species.

Javier Colmenares
- 2022-present – Ph.D. Student Texas Tech
- 2019 – M.S. Universidad Industrial de Santander
- 2015 – B.S. Universidad Industrial de Santander
Contact : jacolmen (at) ttu.edu
My academic and research interests are focused on taxonomy, systematics, and evolution in some groups of small mammals, especially rodents. I’m broadly interested in answering questions about how present and past climatological and geological events influence diversification and population dynamics, and how it is reflected in mammalian genomes. At TTU, I am currently working on a project to understand the evolutionary origin, distinctiveness, and effective population size of a Texas endemic rodent (Peromyscus truei comanche).

Swapnil Boyane
- 2023-present – Ph.D. Student Texas Tech
- 2018 – M.S. University of Pune
- 2016 – B.S. University of Pune
Contact : sboyane (at) ttu.edu
Hello, my name is Swapnil Boyane, and I’m from Pune, India. I’ve always been fascinated by insects and their diversity; therefore, I started working on insect taxonomy while studying for my B.S. and M.S. After obtaining my M.S., I worked as a project associate for ATREE in Bangalore, India and started developing interest in molecular approaches. At TTU, I plan to study the genomics of ants, specifically the genus Camponotus. Additionally, I’ll also try to comprehend how Camponotus and their endosymbionts (Blochmannia) co-evolved.

María C. Tocora
- 2023-present – Research Scholar Texas Tech University
- 2020-present – Ph.D. Student University of Toronto
- 2019 – B.Sc. Biology with Honors, National University of Colombia
Contact : mtocoraa (at) ttu.edu
My research interests revolve around enriching our knowledge of Neotropical biodiversity and understanding how symbiotic interactions shape their evolution. By combining fieldwork, molecular work, and computational approaches, my research seeks to clarify how insect-microbe interactions evolved at micro- and macroevolutionary scales across the Galápagos Islands and in Continental South America. I am specifically investigating how microbes evolve in concert with their hosts, and how host-endosymbiont associations mediate host adaptation to environmental heterogeneity.

Ethan F. Gyllenhaal
- 2024-present – Postdoctoral Fellow Texas Tech / Cornell
- 2024 – Ph.D. University of New Mexico
- 2017 – B.S. University of Rochester
Contact : egyllenh (at) ttu.edu
Google Scholar – Personal Website
My research program uses museum specimens, genomic data, and population genomic simulations to understand the impact fine-scale geography has on island evolution. At TTU, I am working in collaboration with the Messer lab at Cornell to understand the impact geography and selection have had on an island contact zone in the Fiji Whistler (Pachycephala vitiensis), and what that tells us about evolution of island birds more broadly.

Lluis Mercade
- 2024-Present – B.S. Texas Tech University
Contact : lmercade (at) ttu.edu
How species evolved to surpass climate and terrain changes has always been an interest of mine since I played Spore, a video game that shaped the interests I have today. At TTU I am helping Ethan Gyllenhaal to research how geologic histories and change in sea level have shaped the hybrid zone in the Fiji Whistler.

Ashley Allison
- 2022-Present – B.S. Biology with Honors, Texas Tech University
Contact : all14820 (at) ttu.edu
Hi, my name is Ashley and I’m currently working on a project investigating the genomics of the symbiosis and co-evolution of the ant species Camponotus texanus and its bacterial endosymbiont Blochmannia. Ever since my first year of college here, I thought it was interesting to learn that there was a process where a previously free-living prokaryote could evolve & live inside of a eukaryotic organism’s body/cells. This area is of special interest to me because I’d like to study more about the effects that symbiosis has on fitness, reproduction, and genome diversification within both organisms.

Kole Menendez
- 2023-Present – B.S. Texas Tech University
Contact : kmenende (at) ttu.edu
My name is Kole Menendez, I am currently a second-year undergraduate student at Texas Tech University. I have always been passionate about the outdoors and have developed an interest in studying various species, particularly through my experiences in Dr. Manthey’s class. I am currently working on a research project investigating the symbiotic relationship between the Camponotus texanus ant species and its bacterial endosymbiont, Blochmania with others in the lab. Through this research, I aim to gain a deeper understanding of how the fitness and genetic diversity of both organisms is affected, while furthering my knowledge of evolutionary biology and microbial ecology.
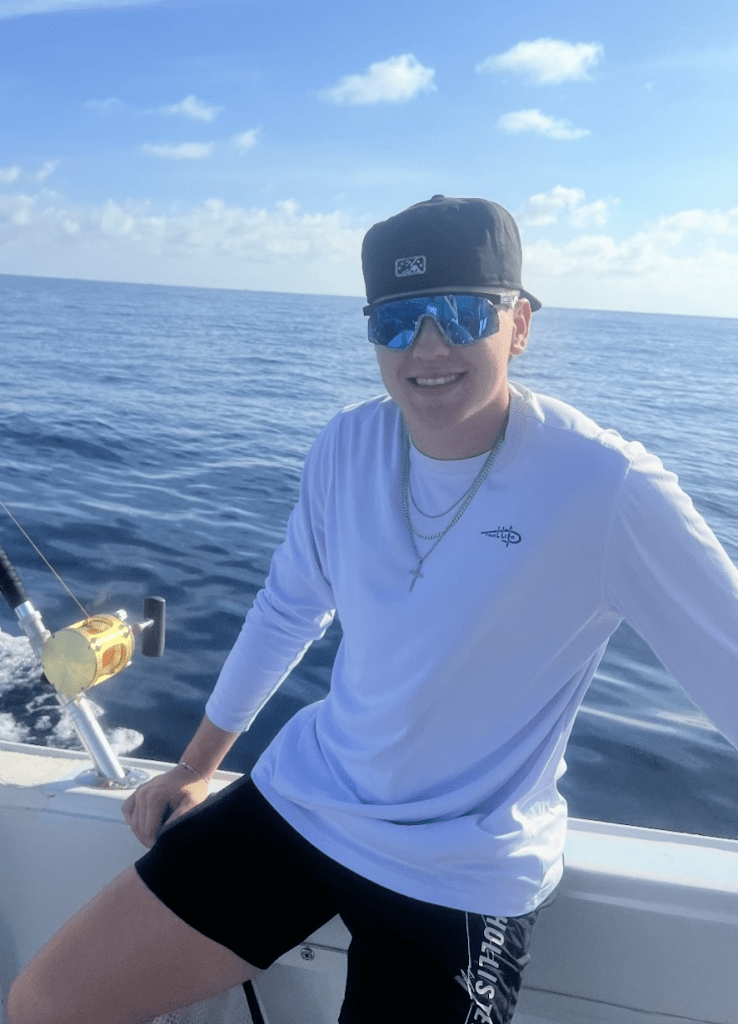

Owen Peck
- 2022-Present – B.S. Texas Tech University
Contact : opeck (at) ttu.edu
Growing up hunting and exploring the outdoors, I developed a fascination with how animals adapt to their environments. At TTU, I am assisting Ethan Gyllenhaal in researching the evolution of island birds, specifically the Fiji Whistler (Pachycephala vitiensis). Our work focuses on the hybrid zone on Vanua Levu, using spectrophotometric data and photographs to quantify plumage patterns across the genus. This research helps us understand how island birds evolve in response to geographic and environmental changes.
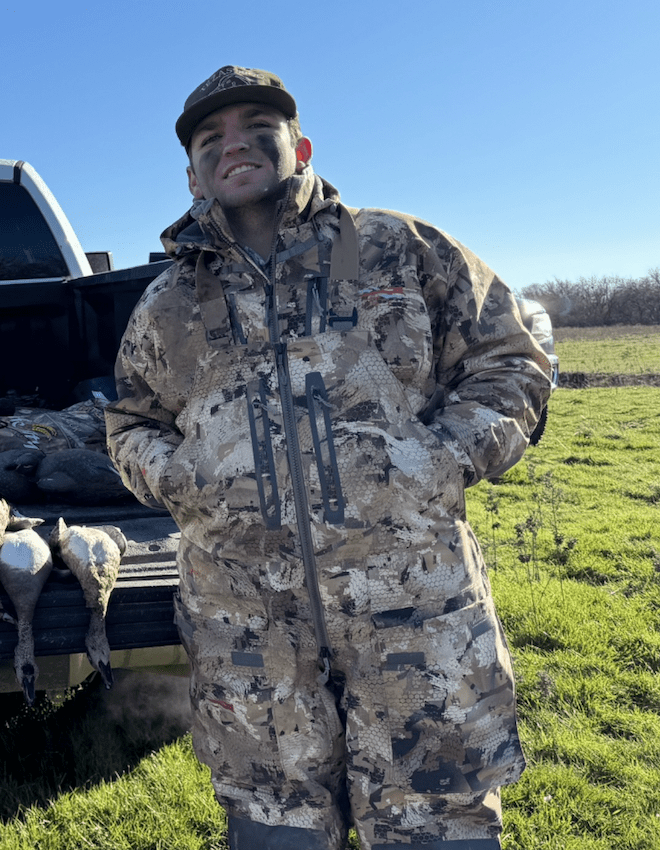

Johanna K. Beam
- 2025-present – Postdoctoral Fellow Colorado State / Texas Tech
- 2025 – Ph.D. The Pennsylvania State University
- 2020 – B.A. University of Colorado, Boulder
Contact : johanna.beam (at) colostate.edu
Google Scholar – Personal Website
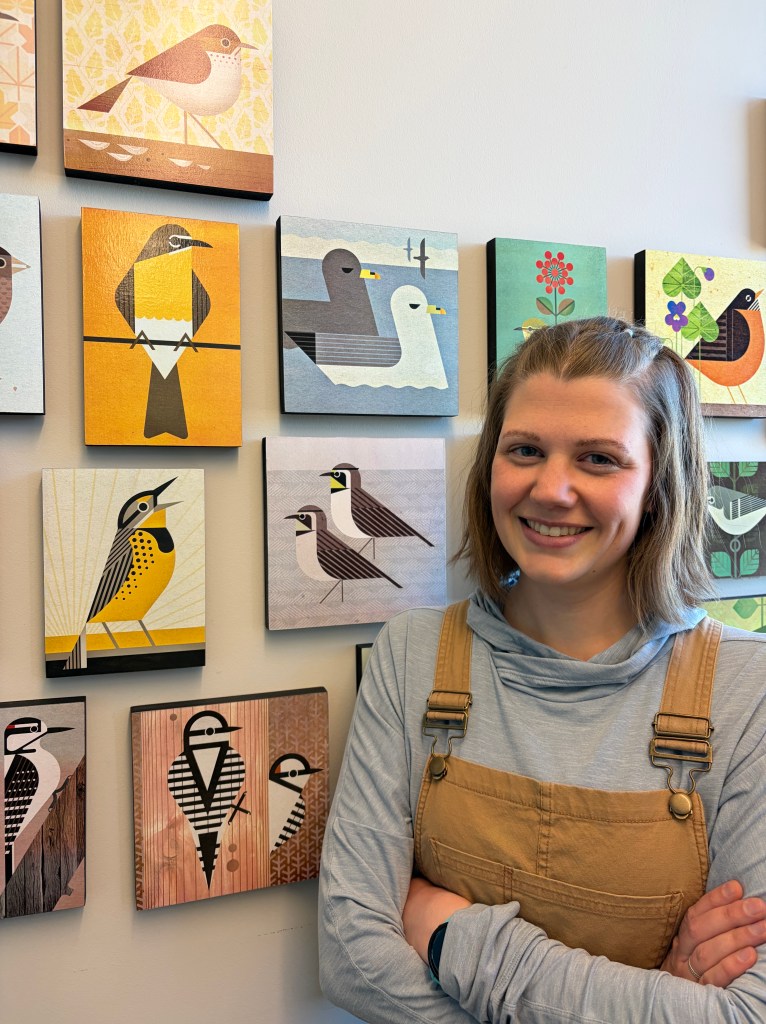
My research focuses on understanding the genomic underpinnings behind birds’ adaptation to their environment and phylogenetic structure. I have previously worked with Sturnella meadowlarks and Yellow-breasted Chats. Now in a collaboration with the Manthey Lab at TTU and the Ruegg Lab / Bird Genoscape Project at Colorado State, I aim to understand how structural variants have shaped the evolutionary history of Horned Lark (Eremophila alpestris species complex). In my spare time, I am also a scientific illustrator and love to do anything outdoors with my dog, Juniper.

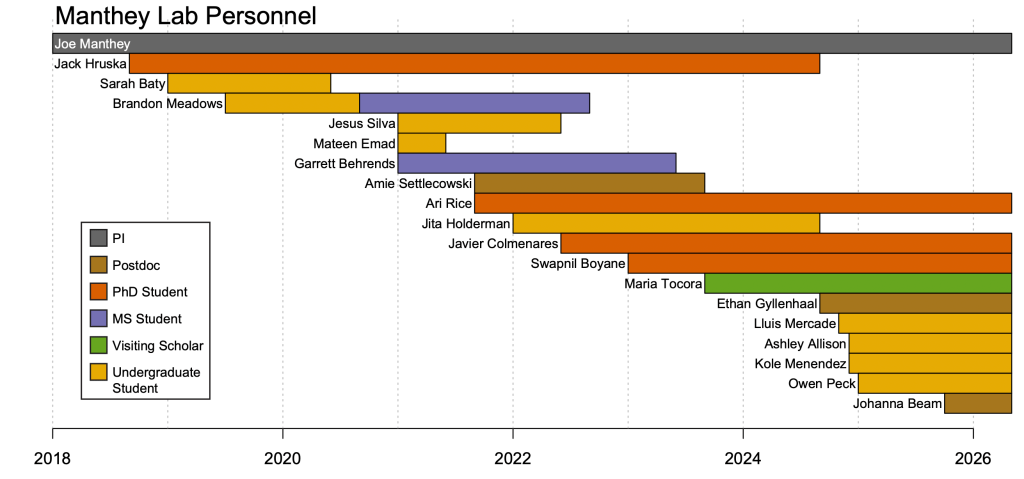
Past Members
Postdoctoral Associates
Amie Settlecowski (2021-2023) | Now Sanger Seq. Manager at University of Chicago DNA Sequencing Facilty
Graduate Students
Jack Hruska (Ph.D. 2024) | Now Postdoc at University of Nebraska - Lincoln
Garrett Behrends (M.S. 2023) | Now Ph.D. student at University of Missouri - St. Louis
Brandon Meadows (M.S. 2022) | Now Ph.D. student at University of Alabama
Undergraduate Students
Alexandra (Jita) Holderman (2022-2024)
Jesus Silva (2021-2022)
Mateen Emad (2021)
Sarah Baty (2019-2020)
Brandon Meadows (2019-2020)